A dawning field of research, artificial biology, is working toward creating a genuinely new organism. At Princeton, chemistry professor Michael Hecht and the researchers in his lab are designing and building proteins that can fold and mimic the chemical processes that sustain life. Their artificial proteins, encoded by synthetic genes, are approximately 100 amino acids long, using an endlessly varying arrangement of 20 amino acids.
Now, Hecht and his colleagues have confirmed that at least one of their new proteins can catalyze biological reactions, meaning that a protein designed entirely from scratch functions in cells as a genuine enzyme.
Enzymes are key to all of biology, Hecht said. “Biology is the system of biochemical reactions and catalysts. Each step has an enzyme that catalyzes it, because otherwise those reactions wouldn’t go fast enough for life to exist. … An enzyme is a protein that is a catalyst. They’re the best catalysts in the universe because evolution has spent billions of years selecting them. Enzymes can increase the speed of a reaction by many orders of magnitude.”
Once Hecht and his research team had successfully created artificial proteins for E. coli, they began looking for critical functions that they could disrupt in these simple bacteria. They found four genes that, when removed, would not only render the E. coli inert — effectively dead — but which their artificial proteins could then “rescue,” or resuscitate.
They first identified these artificial proteins in 2011, and they have spent the past six years working to figure out the precise mechanisms by which their new proteins functioned, now detailed in a Jan. 15 paper in Nature Chemical Biology.
It’s important not to assume that an artificial protein will work the same way as the natural one whose deletion it is rescuing, Hecht cautioned.
After years of experiments, the team had concluded that two of these “rescues” operate by replacing enzymes — proteins that serve to catalyze other reactions, helping them operate quickly enough to sustain life — with proteins that were not enzymes themselves, but which boost the production of other processes in the cell, she said. The third was showing progress, but the fourth had frustrated multiple researchers who came through Hecht’s lab.
But then Donnelly, who was a graduate student when she did the research and is now a research specialist in bioinformatics at the University of Pittsburgh, cracked the code.
“This artificial protein, Syn-F4, was actually an enzyme,” Donnelly said. “That was an incredible and unbelievable moment for me — unbelievable to the point that I didn’t want to say anything until I had repeated it several times.”
She only told Katie Digianantonio, a fellow graduate student, and Grant Murphy, a postdoctoral researcher, who are co-authors of the new paper. “I said, ‘I think this is an enzyme.’ I showed them the initial data and said, ’Don’t say anything to Michael. Let me do this again.” Donnelly re-purified the protein, and created a new, perfectly pure substrate for the E. coli. “I ran everything again from different preps — and when the result held up, I told Michael,” she said.
Out of the original set of proteins that could rescue gene deletions, this is the only one that has turned out to be an enzyme — at least so far, she said.
“We have a completely novel protein that’s capable of sustaining life by actually being an enzyme — and that’s just crazy,” Hecht said.
This has significant implications for industry, said Justin Siegel, faculty director of the Innovation Institute for Food and Health and an assistant professor of chemistry, biochemistry and molecular medicine at the UC Davis Genome Center, who was not involved in the research.
“Biotechnology commonly uses enzymes to carry out industrial processes for the production of materials, food, fuel and medicine,” Siegel said. “The use of these enzymes in an industrial setting often starts with an enzyme that nature evolved for billions of years for an unrelated purpose, and then the protein is tweaked to refine its function for the modern application. The report here demonstrates that we are no longer limited to the proteins produced by nature, and that we can develop proteins — that would normally have taken billions of years to evolve — in a matter of months.”
Hecht’s team had created a strain of E. coli that was missing the enzyme Fes, without which it cannot access the iron needed to sustain life. “We all need iron,” Hecht said. “Even though iron is abundant on earth, biologically accessible iron is not.” Cells have developed molecules like enterobactin, he explained, which can scavenge iron from any available source, but they then need a tool — like Fes — to wrest the iron from the tight grip of the enterobactin.
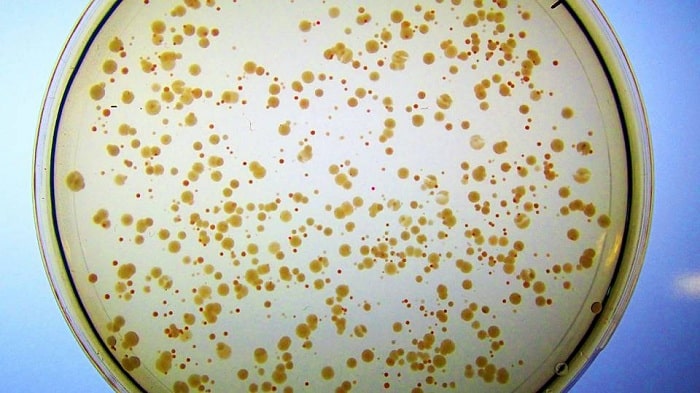
This modified E. coli strain had no way to extract, or hydrolyze, the iron from its enterobactin, until it was “rescued” by Syn-F4. The researchers had provided iron to the E. coli, but it only stained the cells red, since although they could accumulate the bound metal, they could not liberate it from enterobactin or access it for cellular use.
“And then Ann noticed … they aren’t red anymore, they’re white, which suggests the cells can break this down and get the iron, which suggests we actually have an enzyme!” said Hecht.
“Millions of years of evolution resulted in Fes, a perfectly good enzyme for hydrolyzing enterobactin,” said Wayne Patrick, a senior lecturer in biochemistry at the University of Otago in New Zealand, who was not involved in the research. “It is easy enough to study the structure, function and mechanism of Fes, and to infer something about its evolution by comparing it to related sequences. But it is much harder (and more interesting) to ask whether Fes is the solution to the biochemical problem of hydrolyzing enterobactin — or whether it is one of many solutions. Donnelly et al. have shown that an enzyme which was never born (except artificially, in their lab) nevertheless could have been an equally good solution (had it been given the opportunity).
“That line of reasoning has several implications,” explained Patrick. “One is for the life that remains to be discovered on Earth. Perhaps one day, we’ll find a natural enzyme that looks like Syn-F4 but takes the place of Fes in some microorganism or other. At least now, we’ll know to look. Another implication is for astrobiology. If there are many equally likely solutions to a biochemical problem, it becomes more likely that a solution has been found elsewhere in the universe.”
Researchers are on the cusp of a true synthetic biology, Hecht said.
“E. coli has 4,000 different genes,” he said. “We didn’t test all 4,000, because the only way this experiment works is if nothing grows on minimal medium, and of the 4,000, that’s only true for some.
“We’re starting to code for an artificial genome. We’ve rescued 0.1 percent of the E. coli genome. … For now, it’s a weird E. coli with some artificial genes that allow it to grow. Suppose you replace 10 percent or 20 percent. Then it’s not just a weird E. coli with some artificial genes, then you have to say it’s a novel organism.”
This work was funded by the National Science Foundation (grant MCB-1409402) and by the Department of Defense through a NDSEG Fellowship to Donnelly. The authors on the paper, “A de novo enzyme catalyzes a life-sustaining reaction in Escherichia coli,” are Ann Donnelly, Grant Murphy, Katherine Digianantonio and Michael Hecht.